Phiala Thouvenin, Ph.D.'s Portfolio
Coding Projects and Examples
Converting NumPy arrays into gridfiles for GMT and netCDF4
- Simple function to write either a native Generic Mapping Tools (GMT) gridfile or a netCDF-compatible
from utils import *
import numpy as np
import matplotlib.pyplot as plt
import struct
from netCDF4 import Dataset
Generate a simple array of random numbers to save
noise = np.random.random((50,50))
plt.imshow(noise)
<matplotlib.image.AxesImage at 0x7f59e3b968e0>
Save array as either type of grid file
def array_to_grd(filename,
array,
x_vals=None,y_vals=None, #data
x_name=None,y_name=None, #labels
bin_mask=False, # integer 1-0 mask at pixels
netcdf_write=False, # write standard NetCDF grid
native_gmt=False, # write native GMT binary
slice_array=False, spacing=2, # subsample options
xunits = 'x [unitless]', #x-data name
yunits = 'y [unitless]', #z-data name
zunits = 'z [unitless]', #z-data name
title = 'arbitrary data',
cmd = 'Generated by a custom Python function',
remark = 'None'):
'''
Write 2D array to a COORDS-compliant NETCDF4 .grd file,
compatible with GMT. Useful to make binary (0-1) masks to multiply
against GMT-generated .grd files.
Allows for every [spacing]-th point to be sampled, useful for decreasing
filesize if all elements of array aren't needed.
Assumes and image-like coordinate system, where each (x,y) coordinate
is a pixel, unless exact (x,y) coordinates are given [x_vals,y_vals].
Allows for a GMT-native .grd file to be written with custom dimension and
field names, or a modern netCDF4 HDF5 file to be created.
Partially modified from:
https://github.com/usgs/MapIO/blob/master/mapio/gmt.py
TODO:
- figure out array convention for correct plotting direction in GMT
'''
# subsample array, every [spacing] points
if x_vals is None or y_vals is None:
if slice_array:
xcoords = range(array.shape[1])[::spacing]
ycoords = range(array.shape[0])[::spacing]
array = array[::spacing,::spacing]
else:
xcoords = range(array.shape[1])
ycoords = range(array.shape[0])
else:
if slice_array:
xcoords = x_vals[::spacing]
ycoords = y_vals[::spacing]
array = array[::spacing,::spacing]
else:
xcoords = x_vals
ycoords = y_vals
if netcdf_write:
# create and open netcdf file
gmt_file = Dataset(filename, 'w', format='NETCDF4')
# if no alternative dimension names given, set default
if not x_name or not y_name:
x_name='x'
y_name='y'
# dimensions of file, with an unspecified shape for z
gmt_file.createDimension(x_name, array.shape[1])
gmt_file.createDimension(y_name, array.shape[0])
gmt_file.createDimension('z', None)
# variables, with predefined dimensions, according to swapped default GMT
# name conventions for "x" and "y" dimensions
if bin_mask: #integer ones-and-zeros, at each pixel
X = gmt_file.createVariable(x_name, 'i4', (x_name,))
Y = gmt_file.createVariable(y_name, 'i4', (y_name,))
Z = gmt_file.createVariable('z', 'i1', (y_name, x_name))
else: #simple 32-bit float
X = gmt_file.createVariable(x_name, 'f', (x_name,))
Y = gmt_file.createVariable(y_name, 'f', (y_name,))
Z = gmt_file.createVariable('z', 'f', (y_name, x_name))
xcoords = np.float32(xcoords)
ycoords = np.float32(ycoords)
array = np.float32(array)
# data, with x and y pixel locations, and the array with min and max
X[:] = xcoords
Y[:] = ycoords
Z[:,:] = array
Z.actual_range = np.array((np.nanmin(array),np.nanmax(array)))
# close file
gmt_file.close()
# GMT-native format, with custom labels
# see GMT API documentation:
# http://gmt.soest.hawaii.edu/doc/5.4.2/GMT_API.html#gmt-grids
elif native_gmt:
gmt_file = open(filename,'wb')
gmt_file.write(struct.pack('I',array.shape[1])) #ncols
gmt_file.write(struct.pack('I',array.shape[0])) #nrows
gmt_file.write(struct.pack('I',0)) #node offset, 0=gridline, 1=pixel
gmt_file.write(struct.pack('d',np.nanmin(xcoords))) #x-min
gmt_file.write(struct.pack('d',np.nanmax(xcoords))) #y-min
gmt_file.write(struct.pack('d',np.nanmin(ycoords))) #y-min
gmt_file.write(struct.pack('d',np.nanmax(ycoords))) #y-max
gmt_file.write(struct.pack('d',np.nanmin(array))) #z-min
gmt_file.write(struct.pack('d',np.nanmax(array))) #z-max
gmt_file.write(struct.pack('d',np.diff(xcoords).mean())) #dx
gmt_file.write(struct.pack('d',np.diff(ycoords).mean())) #dy
gmt_file.write(struct.pack('d',1)) #z scale factor
gmt_file.write(struct.pack('d',0)) #z offset
# LABELS, cropped to max char length for required GMT header format
xpad = [0 for i in range(0,80-len(xunits))]
ypad = [0 for i in range(0,80-len(yunits))]
zpad = [0 for i in range(0,80-len(zunits))]
tpad = [0 for i in range(0,80-len(title))]
cpad = [0 for i in range(0,320-len(cmd))]
rpad = [0 for i in range(0,160-len(remark))]
xfmt = '%ib' % (80-len(xunits))
yfmt = '%ib' % (80-len(yunits))
zfmt = '%ib' % (80-len(zunits))
tfmt = '%ib' % (80-len(title))
cfmt = '%ib' % (320-len(cmd))
rfmt = '%ib' % (160-len(remark))
gmt_file.write(xunits.encode())
gmt_file.write(struct.pack(xfmt,*xpad))
gmt_file.write(yunits.encode())
gmt_file.write(struct.pack(yfmt,*ypad))
gmt_file.write(zunits.encode())
gmt_file.write(struct.pack(zfmt,*zpad))
gmt_file.write(title.encode())
gmt_file.write(struct.pack(tfmt,*tpad))
gmt_file.write(cmd.encode())
gmt_file.write(struct.pack(cfmt,*cpad))
gmt_file.write(remark.encode())
gmt_file.write(struct.pack(rfmt,*rpad))
nrows,ncols = array.shape
sfmt = '%i%s' % (nrows*ncols,array.dtype.kind)
# write data after flipping, due to how data is written to file and
# read by GMT as a column of data
gmt_file.write(struct.pack(sfmt,*np.flipud(array).flatten()))
gmt_file.close()
else:
raise Exception('Please specify method of gridfile creation.')
Save, load and plot native GMT .grd file
array_to_grd('test.grd',noise,native_gmt=True)
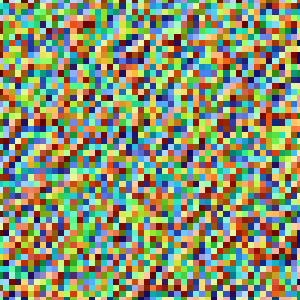
Use the following command in GMT 6 to make a quick plot of the array.
gmt grdimage test.grd -JX1i+ -I+ -png noise

Save, load, and plot a netCDF4 file
array_to_grd('test.nc',noise,netcdf_write=True)
import netCDF4
data = netCDF4.Dataset('test.nc')
noise_retrieved = data['z'][:]
plt.imshow(noise_retrieved)
<matplotlib.image.AxesImage at 0x7f59e329afd0>